Genetic elements promote retention of extrachromosomal DNA in cancer cells
Summary
This Nature study describes a new class of human genomic sequences — called retention elements — that help extrachromosomal DNA (ecDNA) persist in dividing cancer cells. Using a genome-wide functional screen named Retain-seq, live-cell imaging, Hi-C and chromatin profiling, the authors show that retention elements are widespread, enriched at active promoters and enhancers, CpG-rich and hypomethylated. These elements promote tethering of acentric ecDNAs to mitotic chromosomes via contacts with mitotically bookmarked chromosomal sites, increasing nuclear retention across cell divisions. Methylation of retention elements disrupts tethering and leads to loss of ecDNA, and analyses of patient whole-genome data reveal that nearly all oncogene-containing ecDNAs co-amplify retention elements. Evolutionary modelling shows high mitotic retention fidelity is required for selection to amplify ecDNA in tumours.
Key Points
- Retain-seq, a shotgun episome screen, identifies thousands of 1 kb genomic intervals that are preferentially retained as episomes in human cells — termed retention elements.
- Retention elements are enriched at promoters and enhancers, bind bookmarking factors (eg. BRD4, SWI/SNF components), show active chromatin marks and are CpG-dense but rarely act as replication origins.
- Live-cell imaging and Hi-C show retention elements tether ecDNAs to mitotic chromosomes by contacting mitotically bookmarked chromosomal loci, analogous to enhancer–promoter contacts but acting in trans.
- Retention activity is additive: more retention elements on an episome increases mitotic retention fidelity; a single element reduces hitchhike failure from ~25% to ~10% per mitosis.
- Hypomethylation of retention-element CpGs is critical — targeted methylation via CRISPRoff or in vitro CpG methylation abolishes retention and increases untethered ecDNA, leading to ecDNA loss and reduced cell viability.
- Analysis of patient tumour WGS shows ~98% of oncogene-bearing ecDNAs contain retention elements; ecDNA size and structure reflect co-selection for oncogenes plus retention elements.
- Evolutionary simulations indicate a retention fidelity per division of ≳0.9 is required for selection to maintain and amplify ecDNA in growing cancer cell populations.
Content summary
ecDNAs in many cancers carry oncogenes and drive high-level expression but lack centromeres, so their inheritance is risky. The authors propose and test a model where discrete DNA sequences on ecDNA — retention elements — mediate hitchhiking onto mitotic chromosomes, enabling reliable passage into daughter nuclei.
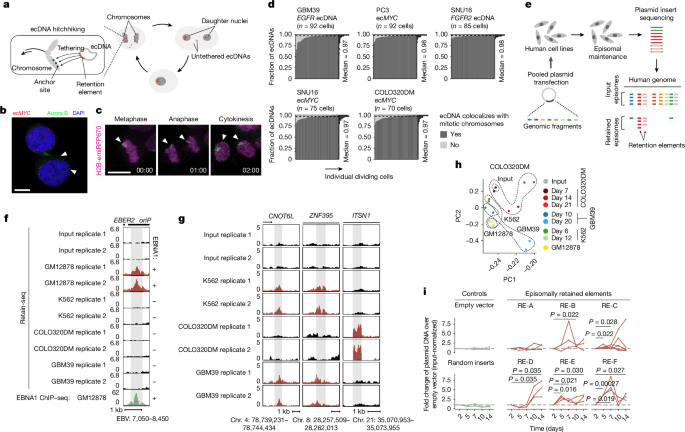
They built Retain-seq by cloning thousands of random human genomic fragments into a simple plasmid backbone, transfecting multiple cell lines and sequencing inserts that persist through serial passaging. The screen recovered known viral tethering sequences (oriP/EBNA1) in the appropriate cell background and revealed >14,000 candidate human retention elements across cell lines. Individual plasmid validations confirmed retention activity for several elements.
Sequence and chromatin analyses show retention elements are promoter/enhancer-like, enriched for active histone marks and bookmarking factor motifs, CpG-rich and often sites of nascent transcription, but they are not primarily origins of replication. Increasing copy number of a retention element on a plasmid boosts retention; conversely, methylation of element CpGs blocks retention.
Live-cell imaging in ecDNA-bearing cancer lines demonstrates that plasmids carrying retention elements colocalise with mitotic chromosomes and are less frequently lost during segregation. Hi-C from mitotic cells shows focal interactions between ecDNA retention sites and mitotically bookmarked chromosomal regions, implying a physical tethering mechanism via bookmarked regulatory sites.
Patient tumour data reveal that almost all ecDNAs that contain oncogenes also include retention elements; larger ecDNAs often carry multiple retention elements, increasing stability. Evolutionary modelling shows that without high retention fidelity, selection alone cannot sustain ecDNA amplification.
Context and relevance
This work explains how acentric oncogene-bearing ecDNAs persist through cell division — a central question for understanding how ecDNAs drive tumour evolution, therapy resistance and intratumour heterogeneity. The identification of retention elements links genomic sequence features and chromatin state (notably hypomethylation and bookmarking-factor binding) to ecDNA maintenance, suggesting new angles to disrupt oncogene amplification. It also provides a generalisable assay (Retain-seq) and mechanistic framework for studying episomal inheritance and designing synthetic episomes for research or therapeutic uses.
Author’s take
Punchy and to the point: this is a big mechanistic advance. The paper doesn’t just label ecDNA behaviour — it pinpoints sequence elements, validates function with imaging and perturbation, and ties the biology back to human tumours. If you care about oncogene amplification, resistance, or genome engineering, read the detail.
Why should I read this?
Want the short version? Researchers finally found the ‘Velcro’ sequences that stick ecDNA to chromosomes. It’s neat, explains why some ecDNAs stick around and others vanish, and reveals a vulnerable epigenetic switch (CpG methylation) you can flip to make ecDNA disappear. If you’re into cancer evolution, drug resistance or synthetic DNA design — this saves you the slog of reading the whole field yourself.