Common variation in meiosis genes shapes human recombination and aneuploidy
Summary
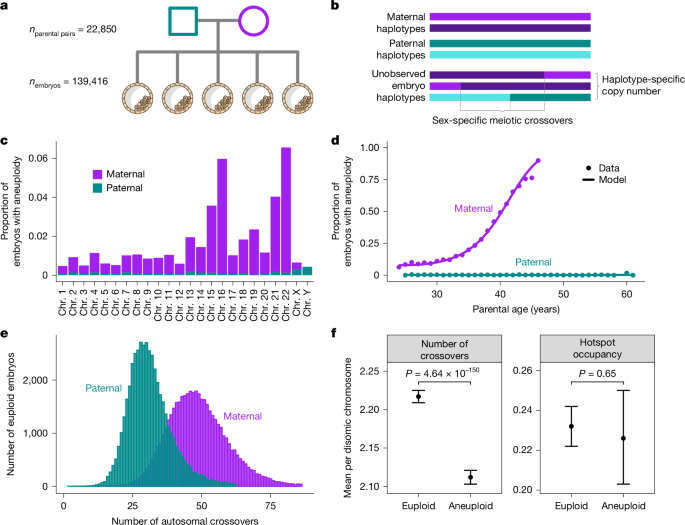
This study used SNP-array pre-implantation genetic testing (PGT) data from 139,416 blastocyst-stage embryo biopsies and genotypes from >22,000 parent pairs to map meiotic crossovers and aneuploidy at scale. The authors developed karyoHMM to trace parental haplotype transmission, call crossovers and detect aneuploid chromosomes. Roughly 29.8% of embryos carried at least one aneuploid chromosome; most aneuploidies were maternal in origin and increased with maternal age.
Analyses show that embryos with meiotic aneuploidies tend to have fewer maternal crossovers on disomic chromosomes, supporting the established idea that missing or poorly placed crossovers raise segregation risk. Genome-wide association and transcriptome-wide tests implicated common regulatory variation at meiosis-related genes. A notable finding is a common haplotype on chromosome 22 (lead SNP rs6006737) spanning SMC1B that associates with higher maternal aneuploidy risk and with reduced SMC1B expression. Fine-mapping points to rs2272804 in a promoter-like element that lowers ATF1 binding in vitro, offering a plausible regulatory mechanism.
Other meiosis genes — including C14orf39 (SIX6OS1), RNF212 and CCNB1IP1 — showed associations with recombination phenotypes and, in some cases, nominal links to aneuploidy. Despite finding reproducible loci, SNP heritability for maternal aneuploidy was low, indicating a substantial role for environmental factors and/or rare variation. Evolutionary modelling suggests that, although the SMC1B risk allele reduces a proxy of reproductive fitness, its persistence at intermediate frequency is consistent with complex relationships between genotype, realised fertility and selection.
Key Points
- Large-scale PGT dataset (139,416 embryos; 22,850 parental pairs) analysed with a bespoke HMM (karyoHMM) to call crossovers and aneuploidies.
- ~29.8% of embryos carried ≥1 aneuploid chromosome; aneuploidies are strongly biased to maternal origin and increase with maternal age.
- Aneuploid embryos show fewer maternal crossovers on disomic chromosomes — supporting the protective role of crossovers against mis-segregation.
- GWAS/TWAS identify common regulatory variation at meiosis genes; a chromosome 22 haplotype (rs6006737) spanning SMC1B is reproducibly associated with higher maternal aneuploidy risk.
- Functional evidence points to rs2272804 (near the SMC1B promoter) reducing ATF1 binding and lowering SMC1B expression — a plausible non-coding mechanism for increased aneuploidy risk.
- Additional meiosis-related loci (C14orf39/SIX6OS1, RNF212, CCNB1IP1) influence recombination and show links to aneuploidy or reproductive ageing traits.
- SNP heritability for maternal meiotic aneuploidy is low, implying major contributions from environment and/or rare variants despite some common-variant signals.
Content summary
The authors combined embryo PGT genotypes and parental data, applied karyoHMM to reconstruct parental haplotype transmission, and catalogued crossovers and aneuploidies at scale. They validated known age and maternal-origin effects and quantified crossover landscapes (median resolution ~99 kb). By comparing disomic chromosomes from aneuploid versus euploid embryos, they showed a clear depletion of crossovers in aneuploid cases.
Genome-wide scans for recombination traits recovered previously reported loci and uncovered links between recombination phenotypes and aneuploidy. A genome-wide significant association for maternal meiotic aneuploidy localised to a common haplotype spanning SMC1B; follow-up eQTL, fine-mapping and EMSA experiments support a regulatory variant (rs2272804) that reduces SMC1B expression by weakening ATF1 binding. TWAS highlighted C14orf39 and other meiosis genes. Despite these signals, aggregate common-variant explained variance for aneuploidy was small.
Context and relevance
This work advances understanding of why human eggs mis-segregate chromosomes — the primary biological cause of pregnancy loss and many neonatal aneuploidies. By leveraging large clinical PGT datasets, the study links common non-coding variation in core meiotic genes to measurable changes in recombination and aneuploidy risk, bridging molecular mechanism (cohesin, synaptonemal complex, crossover control) with population genetics and evolutionary considerations. The findings are relevant to researchers in human genetics, reproductive medicine, IVF/PGT practice, and evolutionary biology, and they help prioritise genes and regulatory sites for sequencing- or functional-focused follow-up.
Author style
Punchy: the paper packs heavyweight sample size, robust methods and functional follow-up into a tight story — common regulatory variation in core meiosis genes can shift recombination patterns and subtly increase embryo aneuploidy risk. If you work on fertility genetics, this is a notable update that points to specific regulatory variants and genes worth following up.
Why should I read this?
Short version — read it if you care about why human pregnancies fail or why eggs go wrong. The team analysed hundreds of thousands of embryo tests, found common gene variants (not just rare mutations) that nudge recombination and raise aneuploidy risk, and even showed a plausible molecular mechanism for one hit. We’ve done the heavy lifting so you don’t have to plough through all the supplements — this is the new big-picture link between common variation, meiosis machinery and fertility outcomes.