Quantum spin resonance in engineered proteins for multimodal sensing
Summary
This paper reports the creation and characterisation of magneto-responsive fluorescent proteins (MFPs) — MagLOV variants — that show large magnetic-field effects (MFEs) and optically detected magnetic resonance (ODMR) at room temperature in living cells. The proteins are derived from the LOV2 domain and use a bound flavin (FMN) to form a spin-correlated radical pair (SCRP) whose singlet–triplet dynamics are modulated by static and radio-frequency magnetic fields. The authors used directed evolution to generate variants (including MagLOV 2 and MagLOV 2 fast) optimised for contrast and response speed, demonstrated single-cell ODMR (≈10% contrast), high per-cell magnetic sensitivity (η0 ≈ 26 μT Hz−1/2), and applications ranging from lock-in amplification and dynamic multiplexing to spatial localisation via an optically detected fluorescence MRI set-up and sensing of local paramagnetic species (gadolinium chelate attenuation of MFE).
Key Points
- Engineered MFPs (MagLOV family) show large MFEs (MagLOV 2: ≈−50% MFE) and room-temperature ODMR detectable in single living cells (≈10% contrast).
- ODMR follows expected electron spin resonance behaviour (fRF ≈ γ̄e B0), confirming spin-1/2 electron transitions tied to a SCRP involving bound FMN.
- Directed evolution produced variants with distinct dynamics (magnitude and saturation rate), enabling orthogonal signatures for multiplexing.
- MagLOV enables lock-in-style detection in noisy, scattering environments; microfluidic tests distinguished weak MagLOV expression from EGFP with balanced accuracy ≈0.99.
- Spatial localisation achieved by combining ODMR with magnetic-field gradients; a custom fluorescence–MRI instrument localised embedded cell bands to millimetre precision.
- MagLOV MFE is attenuated dose-dependently by paramagnetic MRI contrast agent (gadobutrol), demonstrating sensitivity to the local spin environment.
- Advantages over non-biological quantum probes: genetically encodable, expressible in situ, and amenable to sequence-based engineering (rational design or directed evolution).
- Data and code are openly available (DOI and ENA accession provided), supporting reproducibility and further engineering.
Content summary
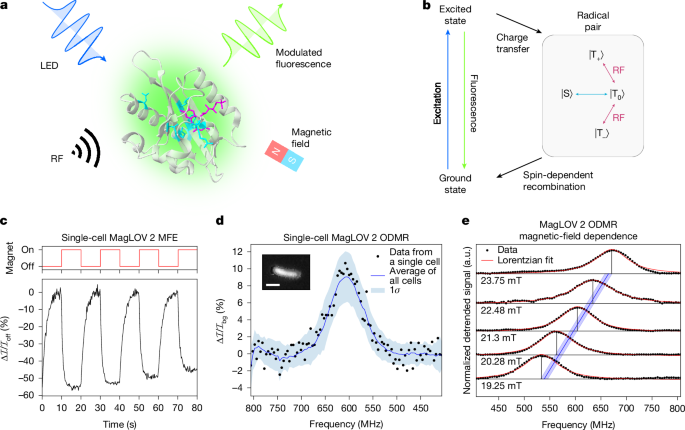
The authors started from an AsLOV2 C450A scaffold and used iterative mutagenesis and fluorescence-based screening to select mutants with enhanced MFE magnitude (MagLOV 2) or faster MFE saturation (MagLOV 2 fast). Using a wide-field fluorescence microscope with applied static and radio-frequency fields, they measured MFEs by switching B0 (0 ↔ 10 mT) and recorded ODMR by sweeping RF at a fixed B0 (≈21.6 mT). Single-cell ODMR signals (≈10% contrast) were extracted and population-averaging improved signal-to-noise across ~1,000 cells.
Spectral and absorption measurements confirm the emission originates from bound FMN. The authors propose a photocycle in which a triplet-born radical pair undergoes singlet–triplet interconversion influenced by hyperfine interactions; application of resonant B1 reconnects triplet sublevels, restoring fluorescence and producing the ODMR signature. Simulations of a proposed photoscheme reproduce MFE and ODMR behaviour.
Practical demonstrations include: (1) dynamic multiplexing — classifying mixed cell populations by differing MFE timescales using an XGBoost classifier; (2) lock-in detection in a mother-machine microfluidic format to reliably pick out weak signals amid autofluorescence/scattering; (3) a fluorescence–MRI proof-of-concept where a radio-frequency coil with a one-dimensional B0 gradient localises multiple MagLOV-labelled cell bands embedded in PDMS; (4) microenvironment sensing via dose-dependent reduction of MFE in presence of gadobutrol, indicating sensitivity to nearby paramagnetic species.
Context and relevance
This work brings quantum-sensing concepts into a genetically encodable, protein-based platform. Until now, most room-temperature quantum sensors suitable for biology relied on non-biological probes (for example, NV centres in nanodiamonds) or required ex vivo conditions. MagLOV proteins combine the advantages of optical fluorescent tags with spin-dependent quantum readout, enabling new measurement modalities: scattering-insensitive localisation, lock-in contrast enhancement, dynamic multiplexing by engineered kinetics, and in situ sensing of magnetic noise sources. The approach aligns with wider trends in quantum-biological probes and optogenetic-style engineering, and could open routes to magnetically controlled biological actuators and in vivo microenvironment sensors.
Author style
Punchy — this is framed as a paradigm shift: genetically encodable quantum sensors that are straightforward to express and engineer. If you care about imaging, biosensing, or synthetic biology, the details matter because the platform directly ties quantum readout to DNA-level programmability.
Why should I read this?
Quick take: they made proteins that behave like tiny, expressible quantum sensors. So if you want better ways to pull signals out of noisy, scattering tissue, do multiplexing with fluorescent tags, or sense local free radicals/paramagnets inside cells — this paper shows the first practical steps. It’s clever, usable and likely to spawn follow-up tools.