A prophage-encoded abortive infection protein preserves host and prophage spread
Article Date: 28 January 2026
Article URL: https://www.nature.com/articles/s41586-025-10070-6
Article Image: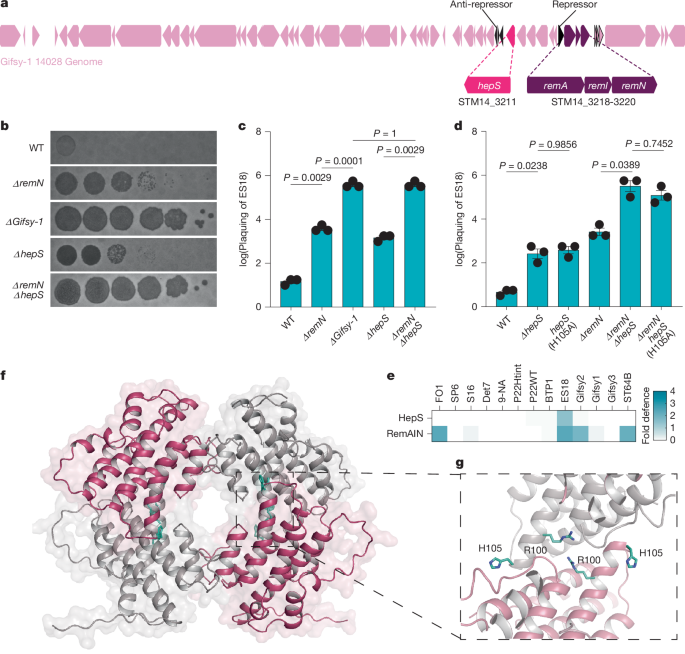
Summary
This Nature study identifies and characterises HepS, a prophage-encoded HEPN-domain abortive infection protein from Salmonella Typhimurium Gifsy-1. HepS is expressed constitutively as an inactive tetramer; it is directly activated by a short peptide from the J tail-tip protein of Siphoviridae phages during phage particle production. Trigger binding remodels HepS into an open, catalytically competent form that acts as an anticodon nuclease, cleaving specific tRNAs (notably tRNA Thr, and to a lesser extent tRNA Ile and tRNA Ser). Activation of HepS aborts phage replication and prevents production of infectious particles, protecting the bacterial host and limiting competing phage/prophage spread. Co-resident Gifsy prophages avoid self-harm by encoding non-triggering variants of the J protein, permitting compatibility between defence and prophage replication.
Key Points
- HepS is a prophage-encoded HEPN-domain abortive infection effector in Salmonella Typhimurium Gifsy-1.
- In the resting state HepS forms an inactive tetramer; binding of the J tail-tip trigger peptide opens the tetramer and repositions catalytic residues.
- Activated HepS is an anticodon nuclease that cleaves specific tRNAs (tRNA Thr most strongly), shutting down protein production and aborting phage assembly.
- HepS is triggered by the J tail-tip protein of Siphoviridae phages; single-residue changes in J (e.g. F924S) generate escape phages that avoid activation.
- Gifsy prophages (Gifsy-1/2/3) express non-triggering J variants (I at the trigger position), allowing these prophages to coexist with HepS while still protecting the bacterial niche from incoming Siphoviridae.
Content summary
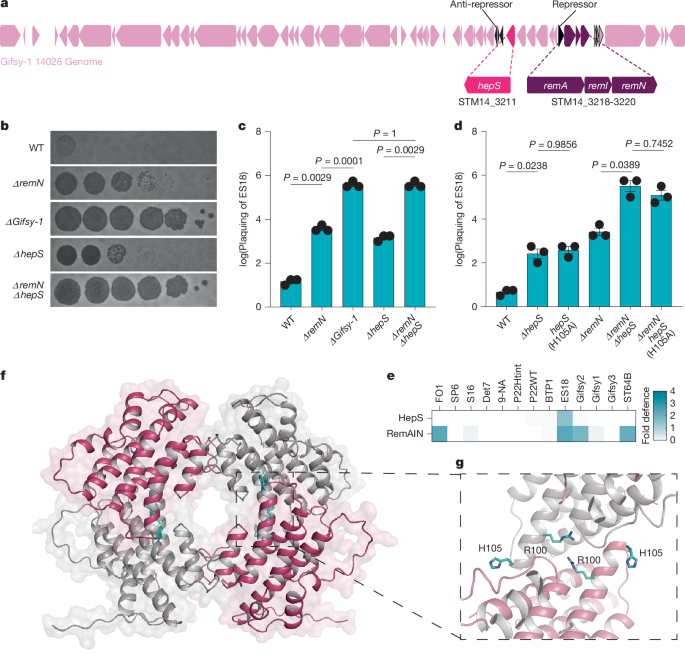
The authors began by searching the Gifsy-1 prophage for defence elements and identified STM14_3211 (HepS) with a canonical RX4-6H HEPN motif. Deletion or catalytic-site mutation of hepS increased susceptibility to representative Siphoviridae phages. Structural biology (2.05 Å apo structure; 1.9 Å trigger co-complex) shows HepS is a tetramer with buried catalytic residues in the apo state; binding of a 32-residue J-region peptide creates a composite β-sheet and allosterically repositions residues to form active HEPN sites.
Functional assays (cellular expression, cell-free systems, tRNA sequencing) demonstrate that trigger-bound HepS cleaves anticodon loops of specific tRNAs (Thr > Ile, Ser), producing tRNA halves and blocking phage protein production/assembly. Phage escape mutants mapped to a single residue in J (F924S) and retain binding but fail to trigger the activating conformational change. In nature, Gifsy prophages carry non-triggering substitutions at the equivalent J position (I rather than F), which prevents self-activation and permits prophage induction and spread while keeping incoming Siphoviridae at bay.
Context and relevance
This work extends our understanding of prophage-encoded defence systems and how they are tuned to avoid autoimmunity. HEPN RNases are emerging as widespread abortive infection effectors; HepS is a clear example of a trigger-activated, highly specific anticodon nuclease that couples structural recognition of a phage tail protein to tRNA cleavage. The study highlights an evolutionary balance: defence that protects the bacterial host and the encoding prophage’s niche, while co-resident prophages evolve subtle changes to their structural proteins to remain compatible.
Implications span basic phage–host interaction biology and applied areas such as phage therapy and synthetic biology. Understanding trigger–effector pairs and escape mechanisms may guide design of phage cocktails or engineered defences and inform predictions about phage resistance evolution.
Why should I read this?
Short version: if you care about how bacteria and prophages fight off invading phages (or how phages dodge those defences), this paper is a neat molecular story. It shows a tidy trigger-and-response system — a prophage-built weapon that only fires when it senses the enemy assembling its tail. The work explains both the biochemistry (structure→activation→tRNA cleavage) and the evolutionary workaround that lets prophages keep their own life cycle intact. We’ve done the heavy reading for you — this is the one to skim if you want the mechanism without wading through dozens of supplementary figures.
Author style
Punchy: the authors combine genetics, structural biology and cell-free assays to deliver a tight mechanistic narrative. The finding is strong for anyone studying bacterial innate defences, phage biology or molecular arms races — it’s not just descriptive, it explains how trigger binding remodels an inactive HEPN tetramer into a precise anticodon nuclease and why that matters for prophage compatibility.