Human and bacterial genetic variation shape oral microbiomes and health
Article Date: 28 January 2026
Article URL: https://www.nature.com/articles/s41586-025-10037-7
Article Image: 
Summary
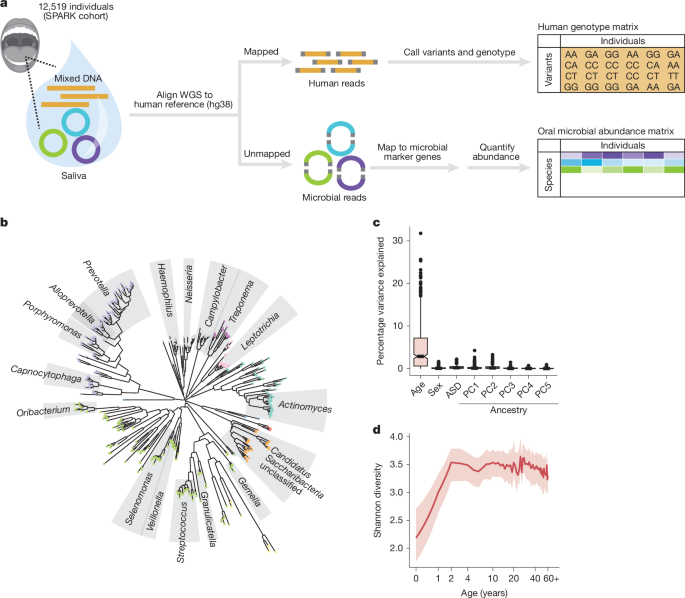
This Nature paper uses whole-genome sequencing (WGS) data from saliva samples of 12,519 people (the SPARK cohort) to link human genetic variation to oral microbiome composition and oral-health outcomes. The authors generated the largest oral microbiome dataset to date (measuring hundreds of species) and developed a principal-component (mPC)-based GWAS to capture pleiotropic host effects on microbial communities. They identify 11 human genomic loci that influence oral microbiome composition, including well-interpretable loci: AMY1 (salivary amylase), FUT2 and ABO (glycosylation/secretor status), TLR1 and HLA class II (immune genes), and PITX1 (tooth development).
Key mechanistic findings include: AMY1 copy-number variation and specific AMY1 missense variants shape abundances of many oral bacteria and associate with dentures/tooth-loss risk; FUT2 secretor status strongly selects for bacterial adhesins and other strain-level gene presence/absence; ABO A1 genotype selects for Prevotella strains carrying a GH95-family glycoside hydrolase that likely uses A antigens as a carbohydrate source. Read-depth analyses across bacterial reference genomes revealed 68 microbial regions (in 18 species) whose gene dosage is associated with host genotypes, suggesting frequent intergenomic adaptation between humans and oral bacteria. Several associations replicate in independent cohorts (UK Biobank, All of Us).
Key Points
- Dataset: saliva WGS from 12,519 individuals yielded the largest oral microbiome profile collection so far (439 species common in ≥10% of participants; 645 species >1% frequency).
- Drivers of variance: host age is a major determinant of oral microbiome composition across the lifespan; ASD status, sex and ancestry explain little.
- Genetics: an mPC-based GWAS found 11 loci affecting oral microbiome composition (P < 5×10−8); many effects are pleiotropic across species rather than limited to single taxa.
- AMY1 effects: salivary amylase (AMY1) copy-number variation (2–32 copies) strongly influences abundances of dozens of species and associates with higher odds of dentures/tooth loss but lower bleeding-gums risk; two AMY1 missense PSVs (F141C, C477R) also greatly increase dentures risk.
- Secretor and blood-group biology: FUT2 non-secretor status (W154X) and ABO A1 genotype strongly shape which bacterial strains and genes persist—e.g., FUT2 selects for bacterial adhesins (YadA-like, CshA, mucin-binding domains) and ABO*A1 enriches Prevotella strains encoding a GH95-like glycoside hydrolase that targets A antigens.
- Microbial gene dosage: read-depth bin analyses identified 208 associations (68 regions) where host genotype correlates with presence/absence or dosage of specific bacterial genes—often strain-level adaptations not reflected in overall species abundance.
- Health links: loci influencing microbiome composition colocalise with dental-health phenotypes (dentures/tooth loss) — especially AMY1, FUT2 and PITX1 — indicating likely causal chains from host genotype → microbiome changes → oral disease risk.
- Methods innovations: the mPC-based GWAS aggregates multivariate microbial signals to boost power and reduces multiple testing; read-depth normalisation and truncated bin phenotypes enable detection of deletions/gene presence in bacterial genomes from human WGS data.
- Replication & robustness: many signals (including microbial gene-dosage effects) replicate in All of Us and UK Biobank samples; careful checks ruled out diet-derived mapping artefacts (e.g., bovine rDNA from dairy).
Context and relevance
This study places oral microbiomes squarely within the host-genetics landscape: compared with gut studies, the oral cavity shows stronger host-genetic influence, perhaps because oral microbes contact host cells more directly. The work bridges human genetics, microbial genomics and oral-health epidemiology and demonstrates molecular mechanisms (glycan targeting, adhesins, amylase-binding proteins) by which host genotype selects for bacterial strains. For researchers and clinicians, the paper offers candidate bacterial drivers of caries and tooth loss and genetic instruments for Mendelian-randomisation follow-up. For anyone interested in personalised medicine, evolutionary genomics or microbiome therapeutics, the findings show how host genotype can shape not just which species are present but which strain-level genes are favoured in an individual’s oral ecosystem.
Why should I read this?
Short version: because it explains how your genes nudge which bacteria live in your mouth — and how that can change your risk of losing teeth. The team read through thousands of genomes so you don’t have to. If you care about oral health, host–microbe interactions or the evolutionary role of diet and glycosylation, this paper is a rich trove of results and methods worth skimming (and worth a deep dive if you work in the field).
Author style
Punchy: big cohort, clear causal candidates, concrete mechanisms. If you’re interested in molecular host–microbe interactions or genetic contributors to dental disease, this is top-tier work — dense but highly actionable. We’ve cut the noise: the highlights above get you to the bits that matter (AMY1, FUT2/ABO, adhesins, GH95 in Prevotella).
Source
Original article: https://www.nature.com/articles/s41586-025-10037-7