Pre-assembly of biomolecular condensate seeds drives RSV replication
Article Date: 28 January 2026
Source URL: https://www.nature.com/articles/s41586-025-10071-5
Image: https://media.springernature.com/lw685/springer-static/image/art%3A10.1038%2Fs41586-025-10071-5/MediaObjects/41586_2025_10071_Fig1_HTML.png
Summary
This study uses single-vRNP live-cell imaging and biochemical tools to show that respiratory syncytial virus (RSV) virions are compositionally heterogeneous and that a minority of incoming ribonucleoprotein complexes (vRNPs) are pre-assembled as condensate ‘seeds’ — called pre-replication centres (PRCs). PRCs carry higher loads of viral phosphoprotein (P), polymerase (L) and nucleoprotein (N), are transcriptionally hyperactive on entry, recruit additional viral proteins and other vRNPs, and reliably nucleate viral factories (VFs). VF formation precedes, and is required for, genome replication; virions that carry PRCs are far more likely to produce successful infections than those that do not.
Key Points
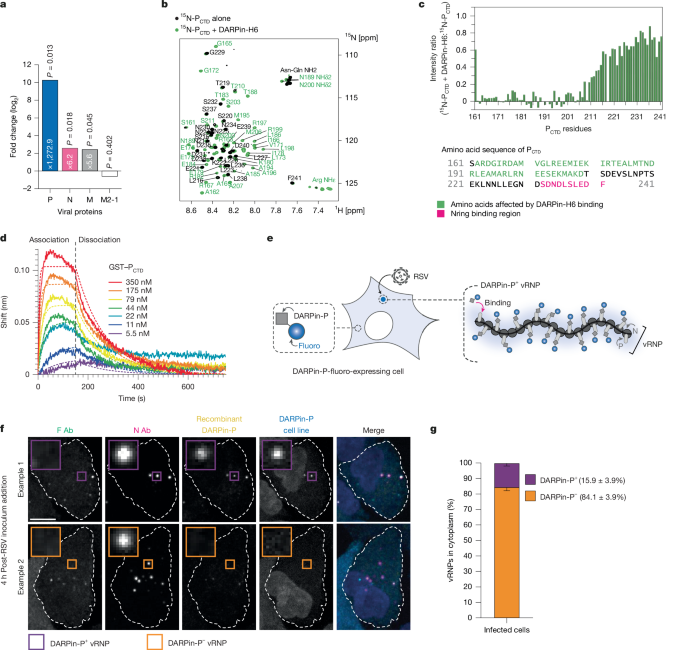
- DARPin-P, a designed ankyrin repeat protein selected to bind RSV P, enables live imaging of a subset of vRNPs and VFs.
- RSV virions contain variable numbers of vRNPs; many virions carry multiple vRNPs that initially co-localise then split after entry.
- A distinct subset of incoming vRNPs (DARPin-P+) — termed PRCs — carry much higher amounts of P and L and are selectively able to seed VFs.
- PRCs show greatly elevated early transcription rates and recruit host‑expressed viral proteins more effectively than ‘passive’ vRNPs.
- VF biogenesis occurs when PRCs grow and fuse with other vRNPs; replication (antigenome production) happens only inside VFs.
- The number of vRNPs per virion is a poor predictor of infection success; the PRC state is the dominant determinant (100% success for DARPin-P+ vRNPs vs ~35% for DARPin-P−).
- PRCs are ‘sticky’ for viral proteins: some of their compositional heterogeneity is packaged in the virion and some is retained after entry because PRCs bind proteins more tightly.
- PRCs behave like condensate seeds and are sensitive to drugs that perturb condensate properties (for example, cyclopamine), suggesting therapeutic angles targeting condensate biophysics.
Content summary
The authors developed imaging strategies to follow individual RSV vRNPs in living cells: DARPin-P (binds P), P exo-fluoro (labels almost all vRNPs via an engineered SunTag/ALFA system), SunTag mRNA reporters (to quantify translation of viral transcripts), and fluorescent tags for L and endogenous P. Using these, they showed that only a subset of incoming vRNPs are DARPin-P positive. Those DARPin-P+ vRNPs (PRCs) have higher P, L and N occupancy, remain more stably protein‑bound after entry, and launch rapid, high-rate transcription immediately after entry.
Timelapse imaging revealed that PRCs grow and fuse with other vRNPs to form large, slow-moving inclusion bodies (viral factories). Antigenome synthesis (replication) is detected exclusively after such VF formation. By contrast, many infections stall: vRNPs enter and split normally but fail to grow into VFs and fail to produce progeny unless a PRC state is reached. Using engineered viruses and single-molecule FISH, the team confirmed that genic RNA integrity is not the main cause of failure; instead, protein composition and dynamic retention/recruitment of P/L/N are decisive.
Context and relevance
This paper addresses a major unresolved point in negative-sense RNA virology and condensate biology: how membrane-less viral factories nucleate early in infection when viral protein concentrations are low. By showing that virions can package pre-assembled condensate seeds (PRCs) that jump-start transcription and nucleate VFs, it explains cell-to-cell variability in infection outcomes and highlights virion composition as a determinant of infectivity.
Relevance to the field: the findings link viral genome/protein packaging heterogeneity to functional outcomes, demonstrate a mechanism that coordinates transcription and replication temporally (transcription first, replication only inside VFs), and suggest that perturbing condensate formation or the stability of PRCs could be a viable antiviral strategy. Because many negative-sense RNA viruses form similar condensates, the result likely generalises beyond RSV.
Author style
Punchy. The authors give clear mechanistic evidence that a small, pre-assembled subset of vRNPs act as seeds for viral condensates, and that these seeds decisively control infection success. For anyone working on viral replication, condensates or early infection dynamics this paper is a crisp, high‑impact read — the microscopy and engineered reporters deliver convincing single-particle resolution data that change how we think about viral factory nucleation.
Why should I read this
Short answer: because it explains why only some RSV particles actually kick off an infection. The team built clever live reporters and watched single viral RNPs in real time — they found a small subset are already primed to make factories and replicate. If you care about how infections start, why outcomes vary between cells, or new antiviral targets (think: condensate physics), this saves you weeks of digging through dense methods — we read it so you don’t have to.