Chemical capture of diazo metabolites reveals biosynthetic hydrazone oxidation
Article Date: 04 February 2026
Article URL: https://www.nature.com/articles/s41586-025-10079-x
Article Image: 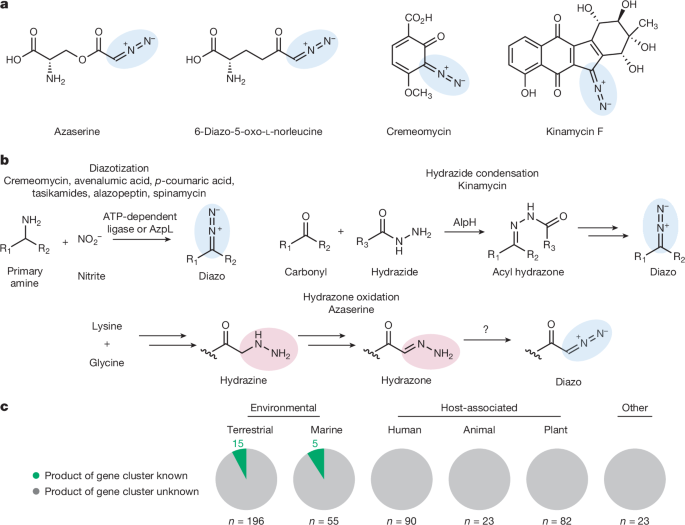
Summary
Researchers developed a reactivity-guided metabolomics workflow that chemically captures labile diazo-containing metabolites using a strained cyclooctyne probe (DBCO-acid). Applying this method to genome-mined bacteria uncovered two previously undetected diazo metabolites—4-diazo-3-oxobutanoic acid and diazoacetone—from Nocardia ninae. The team linked their biosynthesis to a novel dob gene cluster and biochemically characterised Dob3, a ferritin-like diiron oxygenase that catalyses hydrazone N-oxidation to form diazo groups. The work both validates hydrazone oxidation as a diazo-forming strategy and presents Dob3 as a promising starting point for engineering diazo-producing biocatalysts.
Key Points
- Reactivity-based screening with DBCO-acid selectively derivatises diazo groups to stable pyrazole adducts, boosting LC–MS detection sensitivity.
- Genome mining highlighted hundreds of potential diazo biosynthetic gene clusters; the dob cluster in N. ninae produces two novel diazo metabolites (denoted 1 and 2).
- 4-diazo-3-oxobutanoic acid (1) and diazoacetone (2) were only detectable after chemical capture, illustrating how instability masks reactive metabolites in standard screens.
- Heterologous expression of the dob cluster in Streptomyces coelicolor confirmed cluster responsibility for producing the metabolites.
- Dob3 is a ferritin-like diiron enzyme shown in vitro to oxidise hydrazone substrates to diazo products — the first FDO reported to catalyse two-electron N-oxidation of hydrazones.
- Dob3 accepts diverse hydrazone substrates and can form synthetically useful diazo reagents, offering a potential route to enzymatic diazo synthesis for biocatalysis and metabolic engineering.
Content summary
The authors note that diazo functional groups are rare but highly reactive and valuable in both biology and synthesis. Standard discovery pipelines miss many such metabolites because diazo groups are unstable and produced at low titres. To overcome this, the team designed a reactivity-based screen that traps diazo compounds via strain-promoted cycloaddition with DBCO-acid to form stable, MS-friendly pyrazole adducts.
They validated the method with azaserine and applied it to selected bacteria chosen by bioinformatic mining for hydrazone-forming gene clusters. From Nocardia ninae they detected two probe-derived mass features whose formulas matched 4-diazo-3-oxobutanoic acid and diazoacetone. Synthesis of standards (and workarounds for unstable intermediates) confirmed the assignments. Heterologous expression and gene deletion studies linked the dob cluster to the products.
Biochemical reconstitution of the pathway assigned functions to the dob enzymes. Critically, Dob3 (an FDO) oxidises a hydrazone intermediate to form the diazo group; Dob2 (a PKS) performs the C–C bond formation that yields the hydrazone precursor. Dob3 uses a diiron cofactor and molecular O2 and displays activity on multiple hydrazone substrates, revealing scope for enzyme engineering.
Context and relevance
This paper sits at the intersection of natural-product discovery, enzymology and biocatalysis. It shows a practical workflow to reveal chemically reactive metabolites that would otherwise evade detection, and it confirms hydrazone N-oxidation as a genuine biosynthetic route to diazo groups. That has two immediate implications: (1) natural-product diversity hiding in the “reactive metabolite” fraction can be accessed with tailored chemical probes, and (2) enzymes like Dob3 could be evolved as biocatalysts to produce useful diazo reagents for carbene-transfer reactions in greener, biosynthetic settings. The discovery is also notable because the metabolites came from a human-associated pathogen, raising questions about biological roles and possible impacts on virulence.
Why should I read this?
Short version: clever chemistry + genome mining = discovery of previously invisible, highly reactive natural products and a new enzyme that makes diazo groups. If you work on natural-product discovery, enzymatic synthesis, or want biological sources of diazo reagents for biocatalysis, this paper saves you the legwork — they built the probe, proved it works in complex extracts and found both metabolites and the enzyme that makes them.
Author’s take (punchy)
Big win for reactivity-guided discovery. The team didn’t just spot new diazos — they nailed the biosynthesis and pulled out an enzyme (Dob3) with real engineering potential. This isn’t incremental: it opens a route to enzymatic diazo production and extends the known chemistry of ferritin-like diiron oxygenases. Read the methods if you want to copy the screen; read the biochemistry if you want to start engineering Dob3.

