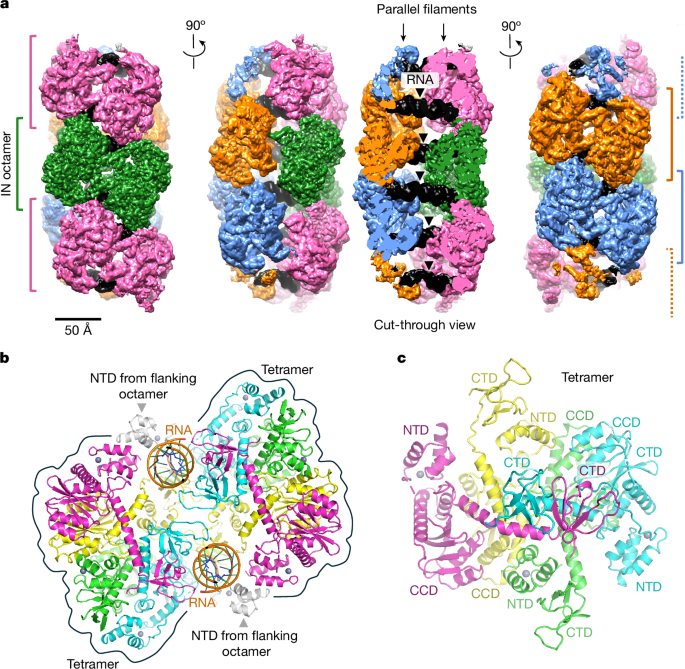
Integrase anchors viral RNA to the HIV-1 capsid interior
Summary
This study shows that HIV-1 integrase (IN) assembles into regular filamentous octameric repeats on the luminal face of the mature capsid and directly captures viral RNA. Using a soluble simian IN (SIVtal) to build IN–RNA co-polymers and high-resolution cryo-EM/cryo-ET of native HIV-1 cores, the authors visualise IN octamer units bridged by RNA duplexes and mapped four IN–CA contact regions on the capsid major homology region (MHR). Mutations in these interfaces, or displacement of IN, lead to eccentric particles that fail reverse transcription. Molecular dynamics, HDX–MS and virology experiments corroborate the structural model and suggest the IN–RNA filament (a ‘proto-intasomal filament’) both staples the RNP to the capsid lumen and prepositions IN for later intasome assembly.
Key Points
- Cryo-EM and cryo-ET reveal IN forms regular filaments of octameric repeats along the capsid lumen that engage viral RNA.
- In vitro SIVtal IN–TAR RNA assemblies provided a 3.3–3.7 Å structure showing CTD-mediated RNA duplex binding and NTD-mediated filament stacking.
- The IN octamer repeat spacing (≈91 Å) and tilt match the mature CA hexamer lattice, allowing direct IN–CA docking via four CA contact regions in the MHR.
- IN or CA mutations at the contact regions yield eccentric virions with RNPs outside the capsid and strong defects in reverse transcription and infectivity.
- IN binding to the CA lattice sterically clashes with LEDGF/p75 IBD binding, implying IN must disengage from the capsid (or be displaced by host factors) to form integration-competent intasomes; ALLINIs target related interfaces and disrupt maturation.
- The authors propose a “proto-intasomal filament”: an IN–RNA assembly that both tethers the genome to the capsid and may seed intasome formation as reverse transcription progresses.
Content summary
The team combined biochemical binding assays (BLI), hydrogen–deuterium exchange MS, single-particle cryo-EM of in vitro SIVtal IN–RNA filaments and cryo-EM/cryo-ET of native HIV-1 cores. The in vitro SIVtal IN–TAR complex produced elongated co-polymers; focused refinement yielded an IN octamer bound to two RNA duplexes. Native core reconstructions (4.6–4.8 Å) and subtomogram averaging (12.6 Å) placed IN octamers on the luminal face of CA hexamers and revealed extra density consistent with the RNP penetrating IN at the RNA-binding sites. MD simulations identified residue-level IN–CA contacts; mutational virology assays validated the functional importance of these interfaces, showing increases in eccentric particles and major defects in reverse transcription and infectivity when disrupted. The work links structural geometry, biochemical binding and functional phenotypes to explain how IN anchors viral RNA during maturation and proposes mechanistic connections to intasome assembly and the action of maturation-targeting inhibitors.
Context and Relevance
This resolves a long-standing question about how the HIV-1 genome is retained inside mature capsids: IN is not just an integration enzyme but also a structural organiser during morphogenesis. The structure explains many previously observed class II IN mutant phenotypes and shows how capsid geometry and IN multimerisation co-operate to staple RNPs to the core. For researchers working on HIV maturation, capsid biology, integrase inhibitors or intasome assembly, these findings materially change how we think about IN function and drug mechanisms.
Author style
Punchy: This is high-impact structural biology — it gives a concrete mechanism tying integrase multimerisation, RNA binding and capsid architecture to viral maturation and early replication steps. If you work on HIV structure, antivirals or RNA–protein assemblies, the full paper is worth a careful read; the supplementary data and deposited maps/models are particularly useful.
Why should I read this?
Quick and blunt — if you care about how HIV packages its genome or how integrase inhibitors mess with particle maturation, read this. The authors have done the legwork: cryo-EM, cryo-ET, MD, mutagenesis and virology all point to the same mechanism. We’ve saved you time — the paper bundles the structural evidence that explains a bunch of weird mutant and drug phenotypes you may have seen elsewhere.