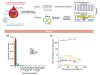
Hidden human–virus interactions uncovered in DNA in blood and saliva
Summary
Researchers scanned DNA sequence data from blood and saliva samples in large biobanks to measure the abundance of viral DNA fragments. They found that viral signals vary systematically with age, sex and dozens of human genetic variants. Notably, genetic analyses indicate that a high abundance of latent Epstein–Barr virus (EBV) in blood is a causal risk factor for Hodgkin’s lymphoma.
Key Points
- Viral DNA fragments detectable in routine sequencing of blood and saliva reveal a human “DNA virome” that differs between people.
- Abundance of viral DNA correlates with demographic factors such as age and sex, and with environmental exposures.
- Genome-wide associations uncovered dozens of human genetic variants linked to variation in viral DNA abundance.
- Mendelian-randomisation and other genetic approaches suggest that higher latent EBV abundance in blood causally increases risk of Hodgkin’s lymphoma.
- Biobank sequencing — even when aimed at human genomes — can yield useful virome data for epidemiology and disease-risk studies.
- Findings point to potential biomarkers and avenues for mechanistic work, but require follow-up to translate into clinical screening or interventions.
Content summary
The study repurposes existing whole-genome sequencing from biobank blood and saliva samples to quantify viral DNA present alongside human DNA. By combining abundance estimates with participants’ age, sex, environmental data and host genotypes, the authors identified systematic patterns in the human DNA virome. Genetic association mapping linked specific human variants to levels of viral DNA, and those variants were used in causal-inference analyses to test whether viral abundance influences disease risk. The strongest causal signal implicated latent Epstein–Barr virus abundance as a risk factor for Hodgkin’s lymphoma, suggesting that viral load — not just exposure — may be relevant for some cancers. The authors emphasise that these results are hypothesis-generating and that mechanistic and clinical validation is needed.
Context and relevance
This work sits at the intersection of genomics, virology and epidemiology. It demonstrates how large-scale human sequencing efforts can be mined to study the virome, offering a new population-level view of host–virus interactions. For researchers and clinicians interested in infection-related cancer risk, immunity and precision public health, the study suggests that latent-virus abundance could be an actionable biomarker. It also feeds into wider trends of using biobank data for multi-omic discovery and for linking genetic variation to infection outcomes.
Why should I read this?
Quick version: if you care about how our genes and environment shape the tiny viral hitchhikers inside us — and what that might mean for cancer risk — this is a neat shortcut. The paper shows you can pull viral signals from ordinary biobank sequencing and that those signals point to a likely causal link between EBV abundance and Hodgkin’s lymphoma. We skimmed the dense bits so you don’t have to — worth a look if you’re into genomics, virology or translational cancer research.
Author style
Punchy: this isn’t just an interesting dataset — it flags a potential causal pathway from latent EBV levels to a specific cancer. If validated, that’s a real translational lead, so read the full paper for methods and implications.