Dissecting gene regulatory networks governing human cortical cell fate
Article Date: 21 January 2026
Article URL: https://www.nature.com/articles/s41586-025-09997-7
Article Image: https://media.springernature.com/lw685/springer-static/image/art%3A10.1038%2Fs41586-025-09997-7/MediaObjects/41586_2025_9997_Fig1_HTML.png
Summary
This Nature study uses a primary human radial glia (RG) culture system combined with single-cell CRISPR interference Perturb-seq and lineage tracing to map how 44 transcription factors (TFs) control cortical progenitor self-renewal, neuronal/glial fate choices, developmental tempo and interneuron (IN) subtype specification. Key findings include discovery of ZNF219 as a regulator that represses neuronal differentiation, opposing roles for NR2E1 and ARX in balancing excitatory versus inhibitory neuron production and timing of lineage progression, and ARX’s post-mitotic role in safeguarding IN subtype identity by repressing LMO1. The work also identifies convergent downstream effector genes enriched for neurodevelopmental and psychiatric disorder associations and demonstrates conservation of many regulatory responses in rhesus macaque.
Key Points
- The authors built a primary 2D human RG differentiation model with high fidelity to in vivo development and lower cellular stress than many organoid/iPSC systems.
- They performed pooled single-cell CRISPRi Perturb-seq targeting 44 TFs (164 sgRNAs) and captured transcriptional and cell-composition effects in >118k cells.
- ZNF219 was identified as a previously underappreciated repressor of neuronal differentiation; NR2E1 and ARX have opposing effects on the excitatory/inhibitory output and developmental tempo of RG.
- Coupling perturbations with barcoded lineage tracing revealed that TF perturbations change fate biases of individual RG clones (EN-biased, IN-biased, mixed), demonstrating lineage plasticity and a restricted developmental window for fate switching (around GW18–19).
- ARX loss produced an ectopic IN subtype marked by LMO1/RIC3; double knockdown of ARX and LMO1 partially rescued that ectopic state, showing ARX represses LMO1 to maintain normal IN identity.
- Convergent differentially expressed genes downstream of multiple TFs are enriched for genes linked to schizophrenia, depression and other neurodevelopmental disorders, highlighting disease-relevant hub effectors.
- Parallel screens in rhesus macaque show broad conservation of regulatory effects, with some species-specific gene-network differences.
Content summary
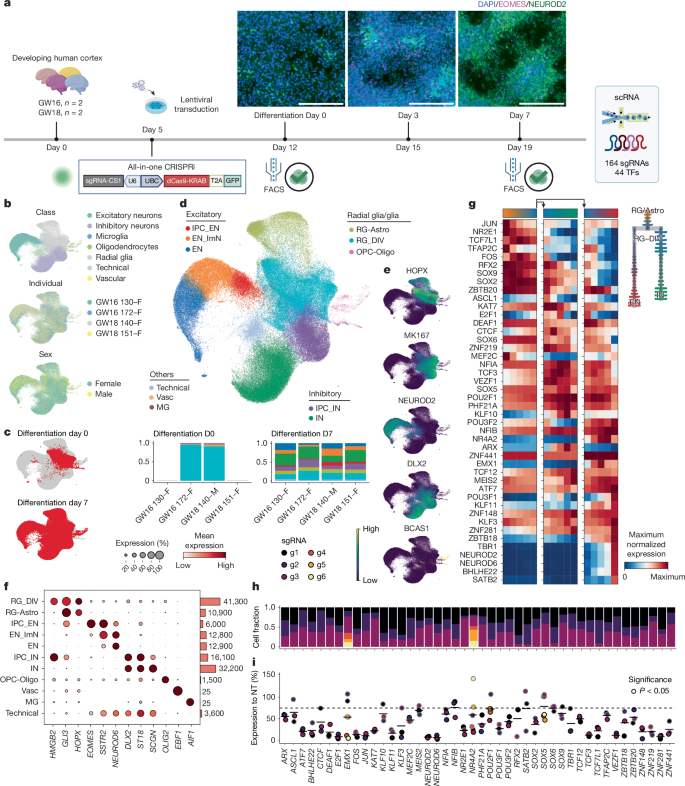
The team isolated primary human radial glia from midgestation cortex, expanded them briefly with EGF/FGF2, delivered an all‑in‑one CRISPRi library (dCas9-KRAB with sgRNA capture) and induced differentiation with BDNF. Single-cell RNA-seq and direct sgRNA capture on a 10x platform assigned perturbations to cells, and KD efficiencies were assessed; inactive sgRNAs were filtered. By day 7 of differentiation, major trajectories (excitatory neurons, inhibitory neurons, oligodendrocytes, astrocyte continuum) were evident.
Systematic TF repression produced graded changes in gene expression and cell-type abundances. The strongest TF phenotypes came from NR2E1, ARX, ZNF219, SOX2, SOX9, CTCF, NEUROD2 and PHF21A. Many differentially expressed genes were shared across multiple TF perturbations and enriched for synaptic, adhesion and neurodevelopmental disease gene sets.
Using cluster-free differential-abundance testing (Milo) and flow cytometry validation, perturbations produced specific composition shifts. NR2E1 KD favoured IN output and accelerated lineage progression; ARX KD favoured EN output and delayed progression. Lineage tracing with STICR barcodes revealed that changes reflect altered clonal fate biases rather than only post-mitotic selection. Pseudotime and clonal analyses mapped heterochronic effects and nominated downstream effector programmes (GLI3/EGFR/PDGFRA axis among others) that distinguish EN- versus IN-producing clones.
Focusing on interneurons, ARX KD generated an ectopic LMO1/RIC3-expressing IN state (conserved in macaque and slice culture) linked to changes in TGFβ/activin signalling, migration and adhesion genes. Double KD of ARX and LMO1 partially restored normal IN transcriptional programmes, supporting a model where ARX safeguards IN subtype identity by repressing LMO1.
Context and relevance
This study provides a scalable functional map connecting TFs to cell fate choices in a physiologically relevant human primary-cell model. It advances understanding of how gene regulatory networks orchestrate the prolonged and complex neurogenesis that underpins human cortical expansion. The work links basic developmental mechanisms to genes implicated in neurodevelopmental and psychiatric disorders, offering candidate effector genes and pathways for follow-up mechanistic or disease-model studies. The combination of CRISPRi (lower toxicity than Cas9 cuts), single-cell readouts and lineage tracing is a powerful blueprint for future functional atlases of human neurodevelopment.
Author style
Punchy: This is a heavy-hitting, method-forward paper — it not only maps who’s in charge (TFs) but shows how flipping those switches rewires fate, timing and subtype identity in human cortical progenitors. If you care about cortical development, lineage plasticity or the developmental origins of neuropsychiatric risk, the details here matter.
Why should I read this?
Quick and tasty: if you want to know which transcription factors actually move the needle on human cortical progenitors — and why that matters for disease — this paper saves you days of combing through supplementary tables. It gives clear candidates (ARX, NR2E1, ZNF219 and others), shows causal clonal outcomes and ties the changes to genes linked to schizophrenia, autism and depression. Proper reading will give you both methods and specific gene leads to act on.