Baby-to-baby strain transmission shapes the developing gut microbiome
Article Date: 21 January 2026
Article URL: https://www.nature.com/articles/s41586-025-09983-z
Article Image: Figure 1
Summary
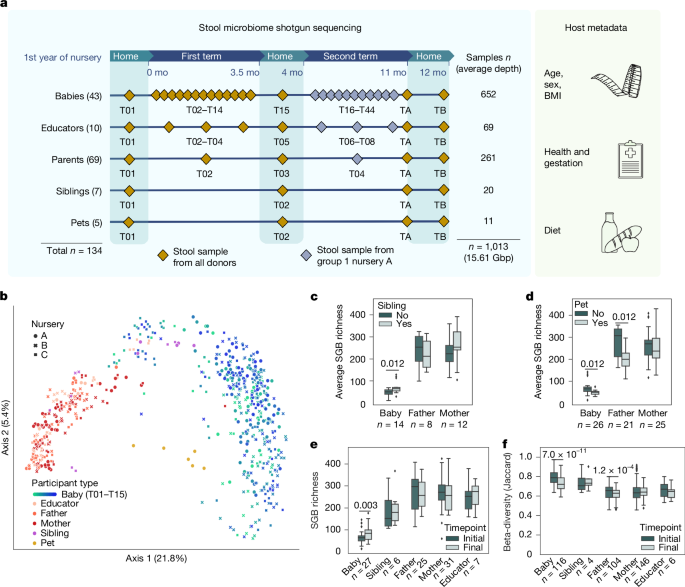
This Nature study (microTOUCH-baby) used dense, strain-resolved metagenomics to track how the gut microbiome of infants changes after starting nursery. The cohort included 43 babies (median 10 months at nursery admission), their families, educators and a few pets (134 volunteers total). Weekly stool sampling of babies across a school term (and follow-ups) plus strain-level profiling with MetaPhlAn4 and StrainPhlAn4 allowed the authors to infer who shared which bacterial strains with whom.
Over the first term babies increased species richness and converged compositionally. Crucially, baby-to-baby strain transmission in the nursery rose quickly: within a month groups showed significantly higher strain-sharing than between different nurseries, and by the end of the term nursery-derived strains made up a larger proportion of infant gut strains than family-derived ones. Some strains spread through a group like outbreaks (for example an Akkermansia muciniphila strain), while Bifidobacterium species (including B. breve and B. longum subsp. infantis) were among the most transmissible and important early colonisers. Antibiotic use strongly reduced strain retention and—particularly in babies—prompted rapid recolonisation via acquisition of new strains. Siblings altered patterns too: babies with siblings had higher baseline diversity and acquired fewer new nursery strains.
Author style
Punchy: this is a tightly designed, high-resolution study that shifts how we think about microbiome acquisition in infancy. It shows social contact among peers—not just parents—actively shapes the infant gut at the strain level over weeks to months. If you care about early-life microbiome assembly, paediatric microbiology or the downstream health implications, the detailed results and methods here are worth digging into.
Key Points
- microTOUCH-baby tracked 1,013 metagenomes (weekly baby sampling) to detect strain-level transmission in nurseries and households.
- Baby–baby strain-sharing rose rapidly after nursery entry; by the end of the first term a larger fraction of infant strains came from nursery peers than from family members (group contribution ≈39.6% vs family ≈20.6% at T15).
- Certain strains spread efficiently within a nursery, sometimes reaching ≥50% prevalence (examples: Streptococcus gallolyticus, Bifidobacterium pseudocatenulatum, Akkermansia muciniphila outbreak chain).
- Bifidobacteria (B. breve, B. bifidum, B. longum subsp. infantis) and other early-life colonisers were highly transmissible among babies—important because these taxa are linked with beneficial functions in infants.
- Antibiotic courses markedly lowered strain retention in both adults and babies but triggered much greater post-antibiotic strain acquisition and strain replacement in babies than adults.
- Having siblings influenced transmission: sibling-associated sharing was high and babies with siblings acquired fewer new strains from nursery peers (possibly due to earlier saturation of strain space).
- Pets showed limited but detectable intra-family strain sharing with babies; cross-family pet–human sharing was rare in this small sample.
Content summary
The study enrolled babies entering their first nursery year in Trento, Italy, with stool samples collected weekly from babies and less frequently from parents and educators. Metagenomes were profiled at species-level genome bin (SGB) resolution and strain phylogenies assembled for hundreds of species. The authors computed strain-sharing rates (SSRs) between sample pairs to quantify transmission.
Findings: babies’ microbiomes became richer and more similar to each other within the same nursery group over the term. Strain-level analysis showed clear, rapid baby-to-baby transfer: SSRs within groups rose from near background to substantial values within weeks. The nursery became a dominant source of new strains for babies during the term. Transmission was not indiscriminate: certain species and traits (aerotolerance, spore-formation, prevalence) correlated with higher transmissibility. Antibiotics decreased within-individual strain retention and promoted recolonisation: babies regained diversity quickly but via many newly acquired strains. Having older siblings shaped baseline diversity and changed the balance of nursery-acquired strains.
Context and relevance
Previous work focused on vertical (mother-to-infant) transfer and household transmission. This study provides strain-resolution evidence that horizontal, intra-generational transmission among peers is a major force in early microbiome assembly once babies enter group care. The result has implications for understanding immune development, pathogen carriage versus beneficial colonisers, and how interventions (diet, probiotics, antibiotic stewardship) might interact with social environments. It also highlights nurseries as hubs where both beneficial and opportunistic strains can spread—relevant for public-health and clinical guidance on antibiotic use and infection control, and for strategies to promote healthy microbiome development.
Why should I read this?
Short version: if you want the real scoop on how babies pick up their gut bugs, this paper shows it ain’t just mum. Nurseries drive fast, measurable strain exchange—think of it as social microbiome networking. Read it for the data, the strain-tracking methods, and the clear implications for antibiotics, siblings, and early-life microbial development.