A cross-population compendium of gene–environment interactions
Article meta
Article Date = 28 January 2026
Article URL = https://www.nature.com/articles/s41586-025-10054-6
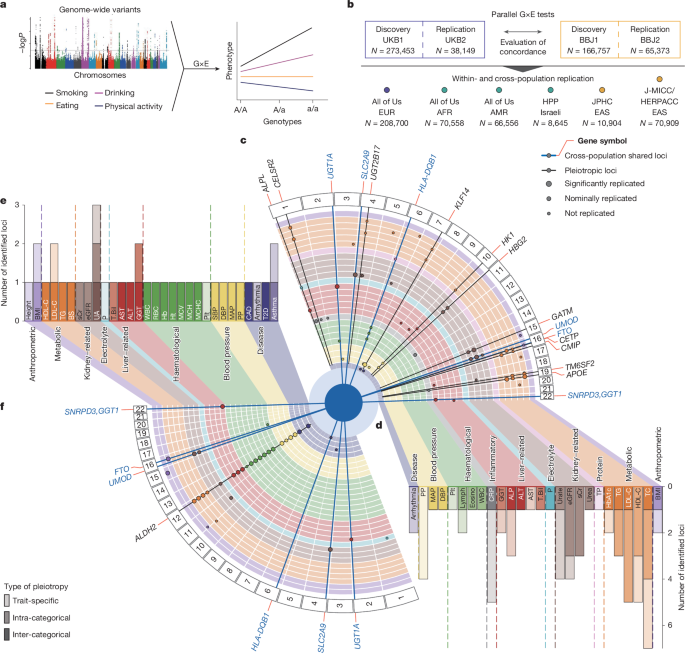
Article Image = Figure 1
Summary
This Nature paper presents a genome-wide, cross-population atlas of gene–environment (G×E) interactions built from large-scale scans in the UK Biobank (European) and Biobank Japan (East Asian). The authors ran parallel discovery and replication analyses, tested 38 biomarkers and 9 diseases against nine environmental factors (age, sex, smoking/drinking, diet and physical-activity clusters), and validated findings in independent cohorts including All of Us and multiple Japanese studies.
They report dozens of genome-wide significant G×E loci (94 trait–locus pairs across 54 loci after meta-analysis), many replicated within and across populations. Key examples include extensive ALDH2-driven, drinking-related effects in Japanese cohorts; environment-driven modulation at FTO, UMOD and ABCG2; and a reverse-causality example linking PITX2, natto intake and atrial fibrillation via warfarin use. The paper further quantifies G×E contributions to heritability, shows how environments alter polygenic score (PGS) accuracy and portability, and integrates omics and single-cell data to connect G×E signals to molecular pathways and cell types (for example, sex-discordant lipid regulation at CETP and an age-shift in pulse-pressure genetics from smooth muscle to endothelial signatures).
Key Points
- Large, parallel G×E scans in UKB and BBJ identified many genome-wide significant interactions, with 64 G×E signals in UKB discovery and 36 in BBJ discovery, culminating in 94 trait–locus pairs across 54 loci after cross-population aggregation.
- Population-specific biology is prominent: ALDH2 (rs671) drives extensive drinking-related pleiotropy in East Asians, while other loci (FTO, UMOD, ABCG2) show environment-dependent effects conserved across populations.
- G×E interactions explain non-trivial fractions of phenotypic variance (G×E heritability), alter polygenic prediction accuracy within and between strata, and can improve prediction when G×E-PGS terms are incorporated.
- Multi-omics and single-cell analyses tie G×E effects to molecular mechanisms and cell types: e.g. sex-discordant metabolite regulation at CETP linked to LDL triglyceride content; age-dependent shift in pulse-pressure genetics from vascular smooth muscle to endothelial signatures.
- Not all detected G×E signals reflect causal gene→environment effects—some reflect reverse causality or healthcare-driven behaviour (PITX2–natto–atrial fibrillation via warfarin), highlighting the need for careful interpretation and temporal analyses.
Why should I read this?
Quick and frank: if you care about why genetic effects change with age, sex, diet or smoking — and why polygenic scores sometimes flop — this paper is a goldmine. It bundles robust, replicated G×E hits across Europeans and Japanese, shows real-world implications for prediction and drug targets, and flags traps such as reverse causality. Read it if you want the big-picture atlas plus concrete locus examples without wading through dozens of separate studies.
Context and relevance
Why it matters: conventional GWAS focus on marginal (fixed) genetic effects and miss plasticity driven by environmental context. This work demonstrates that incorporating environmental context uncovers hidden heritability, explains heterogeneity in risk prediction, and reveals mechanistic detail that can guide drug discovery and trial design (for example, sex-dependent CETP effects that may help explain past trial failures).
Broader implications: the atlas emphasises the need for diverse cohorts with rich environmental and omics data to avoid biased conclusions and to improve equitable genetic prediction. It presents methods and resources (summary statistics, metabolome/proteome QTLs) that researchers and clinicians can use to probe context-dependent genetic effects and to design more precise interventions.