Single-cell and isoform-specific translational profiling of the mouse brain
Summary
This Nature paper presents an advance in single-cell translatomics by adapting Ribo-STAMP (APOBEC1 fused to ribosomal proteins) for use in the mouse brain and combining it with both short- and long-read RNA sequencing. The authors deliver a pipeline that maps translation at single-cell and isoform resolution across the hippocampus, revealing stimulus-induced translation, cell-type-specific translational programmes and isoform-specific ribosome engagement. They also introduce MARINE, a fast edit-calling tool for single-cell edit detection, and apply MAS-ISO-seq for high-throughput long-read isoform sequencing to assign edits to transcript isoforms.
Key Points
- Ribo-STAMP (RPS2–APOBEC1 fusion chosen as optimal) deposits C-to-U edits on translating mRNAs; edits quantified produce an edits-per-reads (EPR) metric that reports translational activity in the same molecules used for transcriptional profiling.
- Ribo-STAMP reliably detects rapid translational changes: BDNF treatment (15′–60′) induced translation of hundreds of genes, many with only modest RNA changes, validated by puromycin–PLA.
- 10x scRNA-seq + Ribo-STAMP in mouse hippocampus yields cell-type-resolved translatomes; EPR patterns identify translation-specific marker genes and show variable RNA–protein coupling across cell types (lowest correlation in oligodendrocytes).
- Single-cell data reveal translational cell states within CA3 and CA1 neurons (bimodal high/low translation groups). High-translation neurons show enriched synaptic and metabolic translation signatures consistent with increased activity.
- Combining Ribo-STAMP with long-read MAS-ISO-seq and MARINE enables isoform-level translation calls (EditsC). Many isoforms of the same gene differ in translation: neuronal high-translation isoforms often have longer 3′UTRs and more nELAVL binding sites, linking isoform choice to translational control.
- Astrocytes and oligodendrocytes display distinct transcript- and isoform-specific translation patterns; oligodendrocytes preferentially show editing on retained-intron isoforms, suggesting cell-type-specific use of intron retention in translational regulation.
- CA3 neurons exhibit higher global translational rates than CA1 (higher EPR, RTE, and AHA incorporation) and increased translation of ribosomal proteins and translation factors; this may relate to basal activity and lower p-eIF2α.
- MARINE (open-source) is introduced as a memory-efficient, fast edit-calling pipeline for single-cell and long-read data; raw and processed datasets are deposited (GEO GSE314176).
Content summary
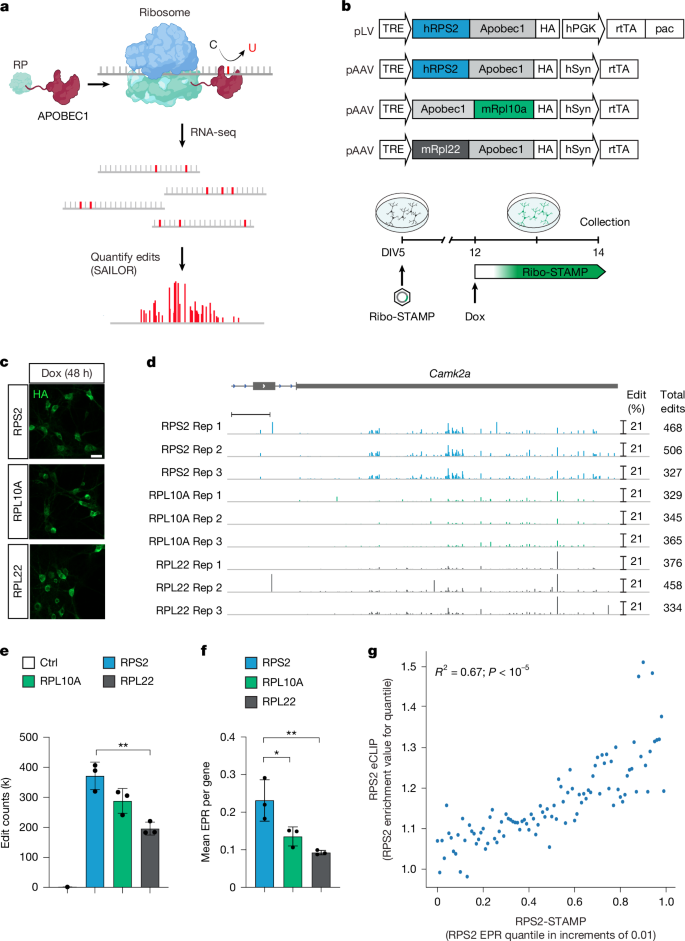
The authors adapt Ribo-STAMP for neuronal expression using an AAV Tet-On system and evaluate variants fused to RPS2, RPL10A and RPL22; RPS2-STAMP performs best and is used for in vivo experiments. They benchmark Ribo-STAMP against eCLIP, polysome profiling and Ribo-seq analogues, and show it tracks translational changes induced by BDNF in primary neurons.
In the mouse hippocampus, constitutive RPS2-STAMP plus 10x scRNA-seq produces ~19,600 high-quality single cells with per-cell EPR values. MARINE identifies barcode-specific C-to-U edits rapidly. Clustering by EPR confirms cell-type-specific translation and yields translation-specific marker genes. CA3 neurons show bimodal translational states; high-translation cells are enriched for synaptic and metabolic translation signatures.
The long-read MAS-ISO-seq data enable isoform assignment of edits (EditsC). Comparing isoforms from the same gene reveals many cases where one isoform is preferentially translated. In neurons, highly translated isoforms tend to have longer 3′UTRs with nELAVL motifs, consistent with positive translational regulation. In glia, motif and RBP analyses implicate RBPs such as PABPC1/4 and SART3 in astrocytic translation preference.
Finally, CA3 vs CA1 comparisons show CA3 has higher global translation and specific upregulation of translation machinery genes; this is supported by FUNCAT, immunohistochemistry and lower basal p-eIF2α in CA3. Limitations include dependence on sequencing depth and editing efficiency; the study provides code (MARINE) and data for community use.
Context and relevance
This work bridges two fast-moving areas: single-cell transcriptomics (including long-read isoform mapping) and translatomics. Previous single-cell translation methods lacked robust isoform sensitivity; by pairing Ribo-STAMP with long-read sequencing and a scalable edit-calling tool, the authors deliver a route to ask which specific transcript isoforms are actually engaged by ribosomes in defined cell types and states. That matters because RNA abundance often poorly predicts protein output in the brain, and isoform choice can dramatically alter localisation, stability and interaction partners.
The paper is directly relevant to researchers studying neuronal plasticity, RNA-binding proteins, alternative splicing, glia biology and neurodegeneration where mis-splicing or translational misregulation is implicated. The datasets and MARINE tool will be useful to labs wanting to profile translation at single-cell and isoform resolution or to validate mechanistic hypotheses about how UTRs and RBPs tune protein synthesis in specific cell types.
Why should I read this?
Short and honest: if you care about what RNAs actually get translated (not just transcribed) in single brain cells — and especially which isoforms of a gene are being used — this paper saves you a ton of guesswork. It gives a practical method, validation, open software (MARINE) and datasets. Read it for the method, the CA1/CA3 translational differences, and the isoform-level findings that would be invisible to conventional scRNA-seq.