Ancient co-option of LTR retrotransposons as yeast centromeres
Summary
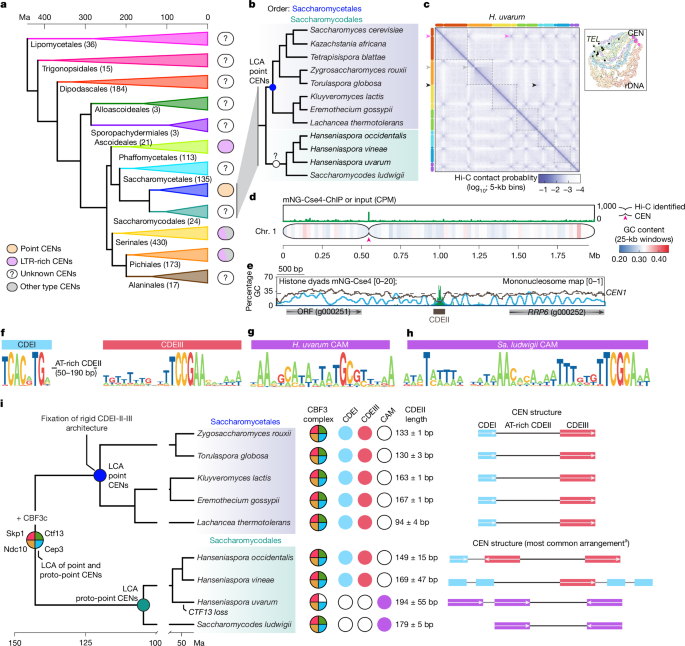
This Nature paper shows that many budding-yeast centromeres originated by co-option of Ty5 LTR retrotransposon sequences. The authors generated chromosome-level assemblies and functional maps (Hi-C, Cse4 ChIP–seq, MNase) for species in the sister order Saccharomycodales and compared them across Saccharomycotina. They identify “proto-point” centromeres: compact, genetically defined loci with a single Cse4 CENP-A nucleosome on an AT-rich core (CDEII-like) flanked by short motif(s) (CAMs) rather than the rigid CDEI–CDEII–CDEIII tripartite structure of Saccharomyces point centromeres.
In Saccharomycodes species these compact cores are embedded in long tracts of Ty5 LTR retrotransposons. LTR age estimates and phylogenetic synteny place Ty5-cluster centromeres deep in the Saccharomycotina lineage. Sequence analyses show that Ty5 LTRs are enriched for binding motifs (Cep3, Cbf1 and GCG-like motifs) and for k-mers typical of CDEII, supporting the idea that Ty5 LTRs supplied the raw DNA motifs that gradually stabilised into CAMs, CDEI and CDEIII. Comparative gene-synteny, protein phylogenies and presence of CBF3 complex subunits argue for common ancestry of proto-point and point centromeres and allow rejection of the 2µ plasmid origin hypothesis. The authors propose a multi-step model: loss of ancestral heterochromatin and Mis18, Ty5 occupation of ancestral centromeric sites, co-option of LTR motifs and progressive evolution of the CBF3–Cse4 system into genetically defined point centromeres.
Key Points
- Discovery of proto-point centromeres in Saccharomycodales: compact AT-rich cores with a single Cse4 nucleosome but variable flanking motifs (CAMs), not the fixed CDEI–CDEII–CDEIII arrangement.
- Saccharomycodes centromeres are embedded in ancient Ty5 LTR retrotransposon clusters; LTR dating indicates long-term occupancy (tens to hundreds of Myr).
- Ty5 LTRs are enriched for transcription-factor motifs (Cep3/Cbf1/Ume6 GCG motifs) and CDEII-like k-mers, providing plausible molecular precursors for centromeric cis-elements.
- Gene synteny and conserved centromere-linked genes across orders indicate Ty5-cluster centromeres were ancestral and inherited by descent rather than repeatedly invented.
- Phylogenetic timing and protein analyses place emergence of CBF3 components before 2µ plasmid invasion, refuting the plasmid-origin model for point centromeres.
- Methods: chromosome-level nanopore+Hi-C assemblies, Hi-C centromere localisation, Cse4–mNeonGreen ChIP–seq, MNase mapping, motif enrichment and LTR phylogenetics across >1,100 genomes.
Content summary
The authors combined new high-quality assemblies and experimental centromere mapping in Hanseniaspora and Saccharomycodes species with broad comparative genomics across Saccharomycotina. Hi-C located compact, non-transcribed centromeric loci; Cse4 ChIP–seq confirmed one Cse4 nucleosome per centromere. In Saccharomycodes, the centromeric cores are flanked and expanded by Ty5 LTR arrays; these LTRs show ages spanning millions to hundreds of millions of years and differ between strains and sister species, indicating ongoing turnover. Motif and k-mer analyses connect centromeric cores and flanking CAMs to Ty5 LTR sequence features; many LTRs carry Cep3 and Cbf1 binding motifs. Parsimony reconstructions and conservation of centromere-linked gene neighbourhoods support an ancestral Ty5-cluster centromere state from which proto-point and, later, strict point centromeres evolved. The work outlines a plausible molecular pathway for the transition from repeat-rich, epigenetic centromeres to genetically encoded point centromeres through progressive co-option of retrotransposon-derived sequences and kinetochore adaptation.
Context and relevance
This study addresses a major question in chromosome biology: how can centromeres become genetically specified when centromere identity is often epigenetic? By demonstrating a widespread, ancient association between Ty5 LTRs and centromeric loci, and by showing LTRs carry motifs recognised by centromere DNA-binding factors, the paper provides mechanistic substance to the idea that transposable elements can seed new, stable centromeric DNAs. The work reshapes models for yeast centromere evolution (rejecting the 2µ plasmid origin) and highlights transposable-element co-option as a driver of functional innovation — a theme with echoes across eukaryotes. For researchers in genomics, evolution and chromosome biology this offers testable hypotheses about motif recruitment, kinetochore evolution and the role of repeat turnover in centromere plasticity.
Why should I read this?
Short answer: because it’s a neat evolutionary detective story with real molecular evidence. If you care about how chromosomes keep behaving while their DNA changes, this paper shows retrotransposons didn’t just crash the party — they provided the furniture. It explains where yeast point centromeres likely came from, uses solid experimental mapping, and gives a clear, testable model for how repeats can be domesticated into essential chromosome features. If you want the mechanisms and timelines rather than hand-wavy ideas, read the details.